
Bovine tuberculosis (bTB) is a disease that affects cattle and, in Great Britain (GB), is caused almost exclusively by the bacterium Mycobacterium bovis (M. bovis). Once almost eradicated in GB, bTB has made a comeback in the last 30 years, particularly after the Foot and Mouth Disease epidemic in 2001.
Bovine TB is an infectious disease, which, similar to human TB causes extensive respiratory disease and eventual death. It is a notifiable disease, and critically, it is zoonotic, meaning it can infect humans. Prior to statutory surveillance introduced in the 1950s, it is estimated bTB caused over 5,000 human deaths annually in the UK – but in many parts of the world it continues to cause human mortality. This risk to humans is one reason we have such strict control measures, and we now very rarely see clinical disease in cattle and have only a handful of human cases per year.
Each year, over 40,000 cattle are compulsorily slaughtered in the UK as part of an effort to eradicate the disease. bTB eradication costs UK taxpayers around £150 million per annum, with additional costs falling to the cattle industry. More information can be found at TB hub - Bovine TB Advice & Tuberculosis Information for Cattle Farmers.
At APHA, over 500 of our staff are involved in combating bTB and our work supports the bTB surveillance and control policies for GB. As part of this, we carry out multidisciplinary research across laboratory, field and data sciences to accelerate innovation and enable the rapid development and adoption of new tools.
In the last few years, we have introduced several new tools and technologies to help in the fight against bTB. Such tools have improved disease detection and monitoring as well as increased disease control. These tools are often initially developed as part of research projects which then make the jump to routine, front-line use following a thorough testing and checking process.
Whole genome sequencing – high resolution DNA fingerprinting of the bacteria which cause bTB
Whole genome sequencing (WGS) is an advanced genetic fingerprinting technique where the entire genetic DNA code (or genome) of an organism (in this case the bacteria which causes bTB, Mycobacterium bovis) is determined. Recent advances in WGS mean that daily analysis of hundreds of different samples is now possible.
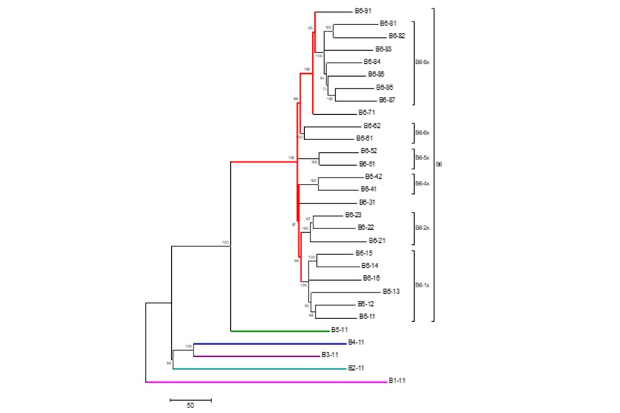
M. bovis can be grouped into different genetic groups called clades, which enable more specific identification of the microbe causing disease in animals, which differ from each other slightly. WGS allows the differences between the individual bacteria to be determined more accurately and with a much higher resolution (ability to distinguish one individual bacterial isolate from another) than previous techniques.
Because different clades of M. bovis are found in distinct geographical regions in the UK, the use of WGS therefore allows movements of different clades of Mycobacterium bovis to be tracked as part of TB outbreaks – in epidemiology investigations.
For example, if a new outbreak occurs on a farm, the sequence of the individual bTB bacteria isolated from the infected cattle can give clues as to where it originated and how it was introduced onto the farm (for example, by looking at and combining the results from cattle movement records).
WGS is also allowing us to investigate important issues, such as the relative contribution of wildlife or cattle movement of bTB spread in defined geographic areas.
Speeding up confirmation of TB in cattle
The standard way to confirm TB in cattle is to take tissue samples from the suspect animal when it is slaughtered and try to grow and isolate the M. bovis bacteria on microbial culture media. Although well-established and widely used, the big disadvantage to this approach is that M. bovis is very slow growing, and it can take up to 22 weeks to get a result. This means farmers can have a long wait to find out if their cattle have bTB.
In an effort to find a quicker solution, we developed a Polymerase Chain Reaction (PCR) test and started using it to test certain bTB cases in March 2022. Everyone is now much more familiar with PCR after the COVID-19 pandemic and the APHA TB PCR test uses the same fundamental approach and principles.
The bTB PCR test detects the presence of M. bovis DNA in bovine tissue samples, even in the presence of very low amounts of genetic material. This can be used as an alternative to culture and takes just a few days. This allows results to be reported and any follow up control measures to be applied more quickly. So far, we have had positive feedback from industry, and we aim to introduce this test for all cases in 2023.
ibTB – allowing everyone to assess the level of bTB risk in their area
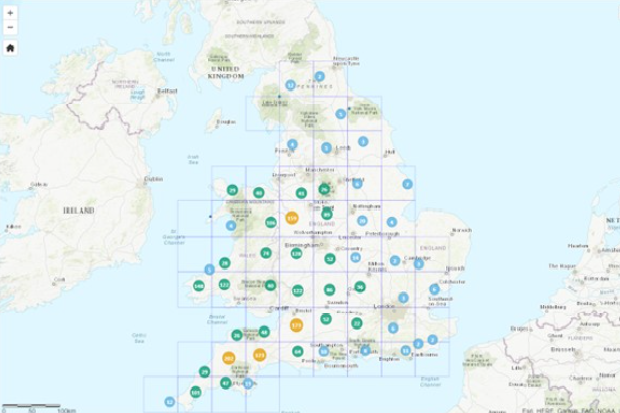
Information bTB (ibTB) is a free-to-access, online interactive mapping tool (developed by APHA and Environmental Research Group Oxford (ERGO)) to help cattle farmers and their vets understand the level of bTB in their area and manage the risks when purchasing cattle. It is based on current and past bTB data and allows users to see the current and historic record of bTB infection in specific geographical locations in England and Wales.
You may have seen mention of ibTB on farming-related shows on television, highlighting its use by farmers and the impact this tool is having to improve their livelihoods.
Since it went live in 2015, the system has proved popular and at the end of 2022 it had its millionth visitor.
Vaccinating cattle against bTB

As with PCR, the COVID-19 pandemic has highlighted the benefits of vaccination to control disease. Vaccination could be an important additional tool in the fight against bTB. So, is vaccination against bTB in cattle possible? We have identified a TB vaccine, CattleBCG, based on the human vaccine bacille Calmette-Guérin (BCG), a weakened strain of the bTB bacteria which does not cause disease. BCG is widely used as a TB vaccine in humans and is the most widely-used vaccine in history, with an incomparable safety record.
So why do we not currently use BCG for vaccinating cattle? Unfortunately, using this vaccine on cows results in cross-reaction with the cornerstone test used in routine surveillance of bTB in cattle – the tuberculin skin test. Any vaccinated animal would give the same positive result as a diseased animal, making it difficult to differentiate between vaccinated and diseased animals.
BCG (just like COVID-19 vaccines) does not give 100% protection, so animals could be vaccinated and still catch the disease. We therefore still need a compatible surveillance test to work alongside vaccination that can Distinguish Infected among Vaccinated Animals – the so-called DIVA test.
In a breakthrough development, APHA researchers have developed a successful DIVA test, based on components of the bTB bacteria which are not present in the vaccine. This companion DIVA test is used as a skin test in a similar way to tuberculin.
In order to bring BCG and DIVA to market and allow routine veterinary use as part of the bTB eradication programme, these new tools are subject to the same scrutiny and application process as any other veterinary vaccine. This involves submitting a dossier of data on the safety and efficacy of the approach to the Veterinary Medicines Directorate and applying for approval. At present, APHA are conducting field trials to allow this data to be collected.
New TB tests for other species
In addition to cattle, Mycobacterium bovis can infect and cause bTB in other domestic and wildlife species such as pigs, deer and camelids. We have recently developed and validated new blood antibody tests that allow bTB testing in these species and provide tools for surveillance and monitoring of the disease in these populations.
Subscribe to our blog
Throughout the year, we publish blogs which highlight the breadth of scientific work we are involved in as well as sharing our latest news and events we have attended.
Subscribing to our blog takes seconds and you will receive instant email alerts as soon as new blogs are published so why not subscribe today!

Recent Comments